Release 1.2
Coot 1.2 Released 1st April 2026
Graphics
-
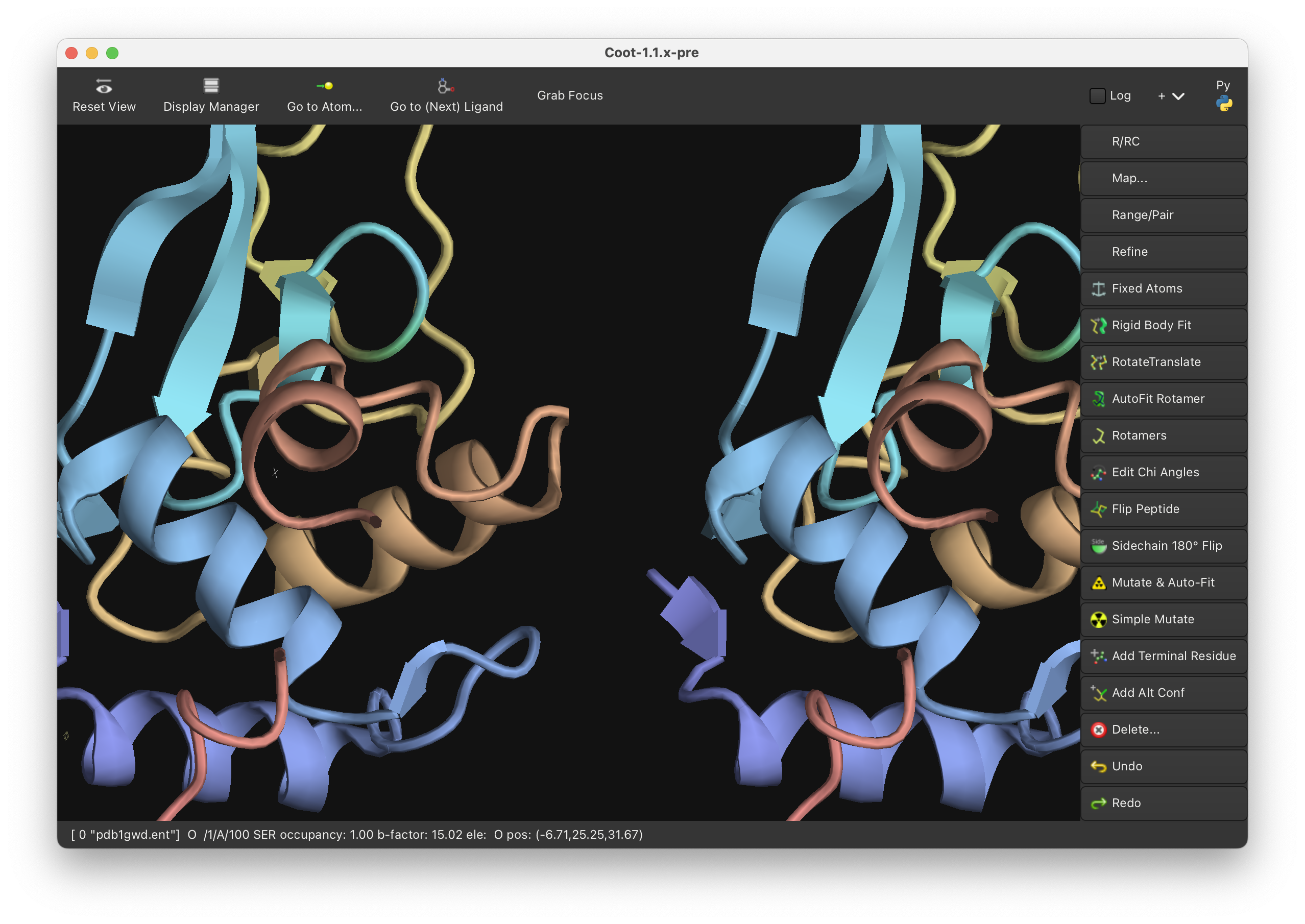
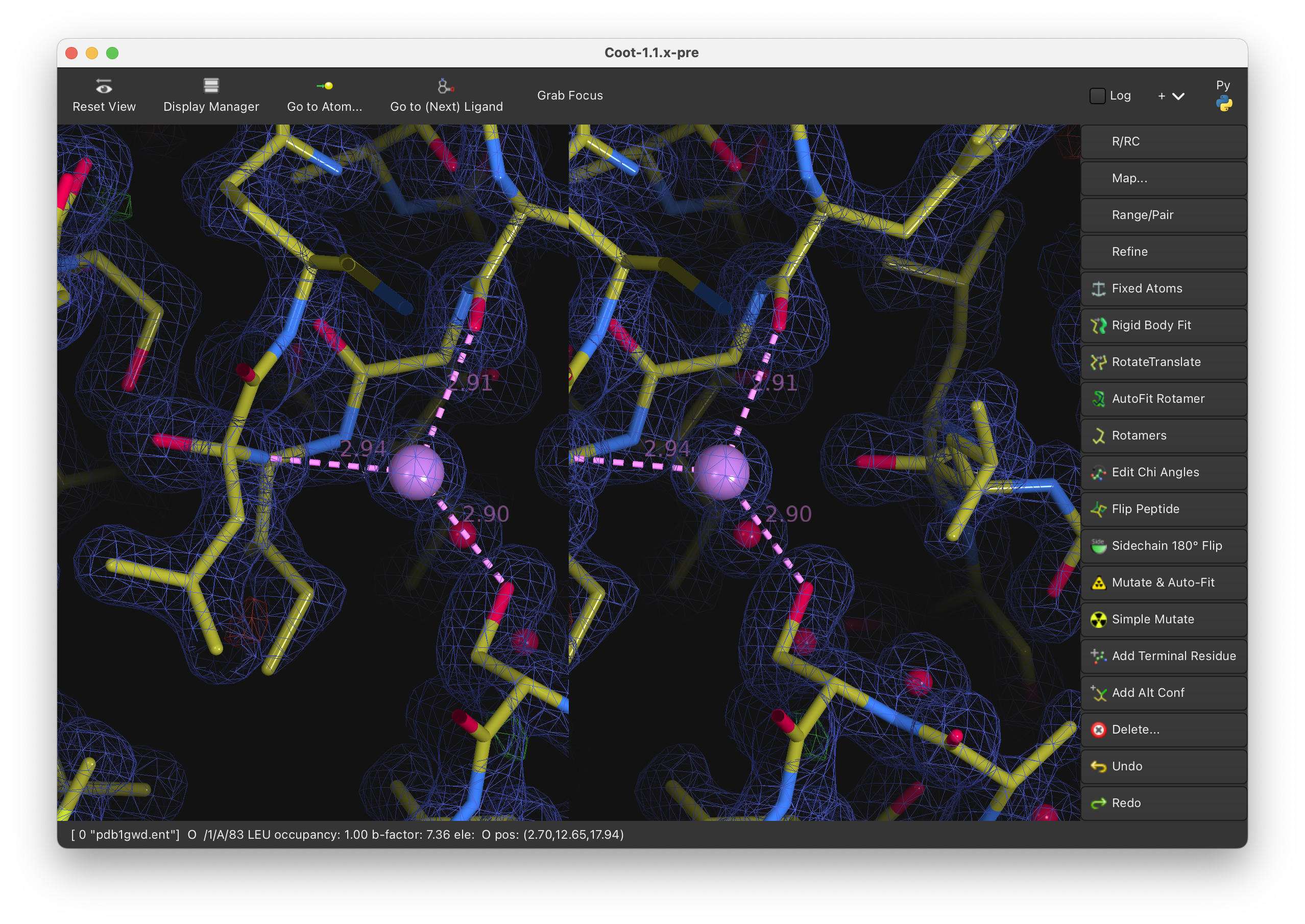
FEATURE: Side-by-side stereo mode (cross-eye and wall-eye) with configurable stereo angle
-
FEATURE: High-resolution screenshots now work for both basic and fancy rendering modes
-
FEATURE: Material properties added to instanced-object shaders
-
FEATURE: Anisotropic thermal ellipsoid “empty” representation with multi-ring display
-
FEATURE: Gaussian surface opacity control
-
FEATURE: Bond smoothness toggle
-
FEATURE: Shadows for “other meshes” (generic display objects)
-
FEATURE: Single-model view for CA+Ligands representation with colour-by-dictionary-and-chain
-
FEATURE: Distance labels display can now be toggled with set_show_distance_labels()
-
FEATURE: scale_up_graphics() and scale_down_graphics() now added to the API
-
CHANGE: Improved default secondary structure colours
-
CHANGE: Rainbow ribbons are less bright
-
CHANGE: Aniso probability scale now goes to 0.99
-
CHANGE: Ribbon coils restored with radius multiplier for spline representation
-
CHANGE: End-caps added for worm representations
-
CHANGE: Max SSAO kernel samples doubled to 512
-
CHANGE: Reduced step size for front clipping plane changes
-
CHANGE: Don’t include hydrogen atoms in Local B-factor display
-
CHANGE: x-ray Sharpen/Blur Map slider now displays the value
-
CHANGE: Difference map peaks now shows closest atom not position
o BUG-FIX: Now the Recover Session works as expected [Kevin Jude]
o BUG-FIX: Fix the rotation jump after panning
o BUG-FIX: Fix peptide straggle-bond when displaying alt-confs
o BUG-FIX: Fix texture indexing in TextureMesh
o BUG-FIX: Fix framebuffer rendering with transparency
o BUG-FIX: Fix the map colour button in single-map properties dialog
o BUG-FIX: Fix ribbon surface normal computation and spline frame orthonormality [Martin Noble]
Modelling
-
FEATURE: Post-Translational Modifications (PTMs) can now be applied to residues, with a database of common PTMs
-
FEATURE: Improved jiggle-fitting for Local Fitting in Cryo-EM
-
FEATURE: Add Residue by Map Fit function
-
FEATURE: Ephemeral overlay labels for transient annotations
-
FEATURE: Key-binding R for Refine with auto-accept
-
FEATURE: Key-binding Ctrl-E for Python scripting entry
-
FEATURE: Mutate-and-auto-fit improvements
o BUG-FIX: Fix self-deadlock in make_moving_atoms_graphics_object() for CA bond mode
o BUG-FIX: Fix single-strand test in ideal nucleic acid builder
o BUG-FIX: Fix PHE/TYR nomenclature error side chain atom swapping
Validation
-
FEATURE: Phenix .geo file validation for bonds, angles, chirals and non-bonded contacts with colour-coded display
-
FEATURE: Validation graphs can now be undocked [Jakub Smulski]
-
FEATURE: Rotamer probability now shown in validation graph
-
FEATURE: Refinement results mini-statistics (bond and angle distortions) returned from refinement
-
FEATURE: Atom geometry distortion info now includes atom specs
-
CHANGE: Molecule mesh display control toggles relocated
o BUG-FIX: Fix rotamer probability in validation graph model list refresh
Ligands
-
FEATURE: Ligand Builder (Lhasa) added to the Ligand menu
-
FEATURE: Fetch Ligand Restraints Dictionary callback added
-
FEATURE: Pyrogen integration via RDKit mol pickle base64 API
-
FEATURE: Curlew extension download and install function
-
CHANGE: Ligand fitting dialog is now scrollable
o BUG-FIX: Fix the bond-order combobox default in acedrg link dialog
o BUG-FIX: Fix Lhasa molecule centering [Jakub Smulski]
o BUG-FIX: Fix RDKit stereochemistry handling and isomeric SMILES output [Jakub Smulski]
Maps
-
FEATURE: NetCDF support for reading AMBER trajectories
-
FEATURE: Slurpable maps recognized as EM maps
-
CHANGE: Improved map-based occlusions
Scripting and Automation/MCP
-
FEATURE: JSON-RPC server, MCP Bridge and Skills for Agentic Coot
-
FEATURE: RPC server menu item added
-
FEATURE: Python API function documentation improvements across the board
-
FEATURE: coot_ping() function for connectivity testing
-
FEATURE: get_function_descriptions() added to the bridge
User Interface
-
FEATURE: Shift-click on an atom shows atom info in the status bar
-
FEATURE: “Delete all water” menu item
-
FEATURE: EM Placement dialog with Phaser integration
-
FEATURE: Git commit hash and build date shown in About dialog
-
FEATURE: Improved “Recover Session” dialog
-
FEATURE: Generic Display Objects dialog now shows button state
-
FEATURE: Interesting places JSON file reading with atom-spec parsing
-
CHANGE: Copy Fragment combobox now sets the active item
-
CHANGE: Don’t include hydrogen atoms in Local B-factor display
-
CHANGE: Sharpen/Blur Map slider now displays the value
-
CHANGE: Difference Map Peaks now shows closest atom
-
BUG-FIX: Fix validation graph combobox g_warning()
-
BUG-FIX: Better handling for unknown monomer types in PTM dialog
-
BUG-FIX: Remove duplicate entries from PTM combobox
-
BUG-FIX: Fix Copy Fragment combobox active item
Chapi/libcootapi
-
NEW FUNCTION: get_molecule_diameter()
-
NEW FUNCTION: get_torsions_for_residues_in_chain()
-
NEW FUNCTION: get_hydrogen_bonds_py() - pythonic hydrogen bond info
-
NEW FUNCTION: set_residue_properties()
-
NEW FUNCTION: get_computed_acedrg_atom_types()
-
NEW FUNCTION: pyrogen_from_rdkit_mol_pickle_base64()
-
NEW FUNCTION: get_rdkit_mol_pickle_base64() with pickling options
-
NEW FUNCTION: add_residue_by_map_fit()
-
NEW FUNCTION: residue_exists_py()
-
NEW FUNCTION: ephemeral_overlay_label()
-
NEW FUNCTION: molecule_atom_overlaps_py() with n_max argument
-
NEW FUNCTION: get_xmap() for testing
-
NEW FUNCTION: get/set view from/to JSON string
-
NEW FUNCTION: set_stereo_angle()
-
NEW FUNCTION: side_by_side_stereo_mode()
-
NEW FUNCTION: curlew_download_and_install_extension()
-
NEW FUNCTION: get_ligand_validation_vs_dictionary() fix
-
NEW FUNCTION: get_molecule_selection_as_json() fix
-
NEW FUNCTION: add_molecular_representation with secondary structure usage flag
-
BUG-FIX: Fix get_residues_in_chain_py()
-
BUG-FIX: Fix user-defined atom colour handling
Build
o Updated RDKit, libsigc++3, Boost, Python versions
o Improved Lhasa build scripts [Jakub Smulski]
o Ubuntu CI workflow improvements
o MAKE_ENHANCED_LIGAND_TOOLS build flag cleanup